Services and Instruments
NEXTGEN SEQUENCING/GENOMICS
The NextGen Sequencing/Genomics core facility at the Center for Biotechnology and Genomics provides sequencing services to the entire TTU community as well as outside customers.
These services include sequencing library preparation from DNA/RNA, sequencing on state-of-the-art platforms, and primary and secondary data analysis. We also standardize new protocols for sequencing library preparation.
A major component of our service includes student training in NextGen sequencing and associated Bioinformatics.
Instrumentation:
- Illumina NovaSeq 6000
- Illumina MiSeq
- 10x Genomics Chromium controller
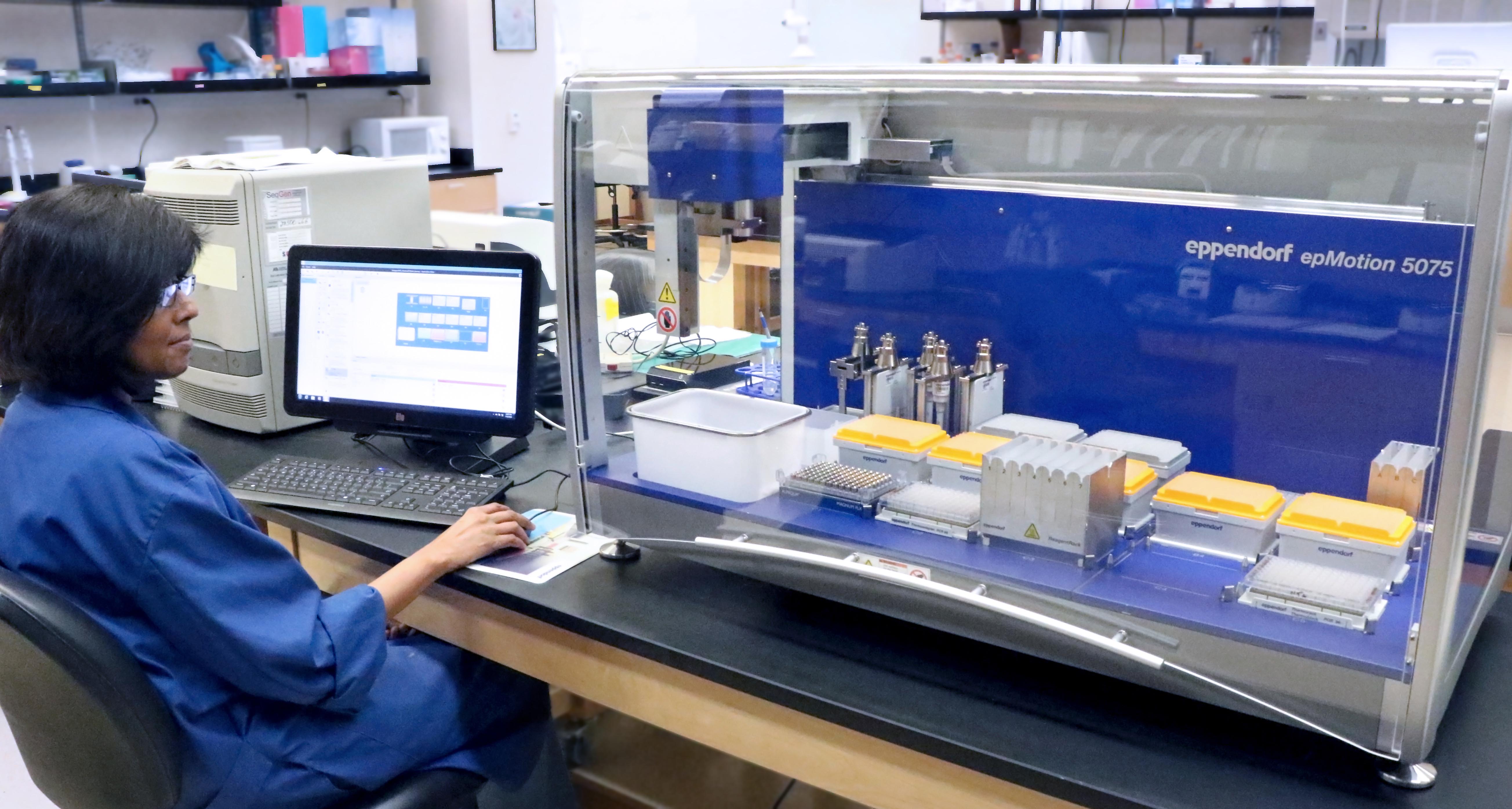
- Eppendorf epMotion liquid handler
- DragonFly and Mosquito liquid handlers
- Biotek plate reader
- Agilent tape stations
- Spectrophotometers and other equipment for sample and library preparations
Services Offered:
- Stranded mRNA-sequencing for both prokaryotes and eukaryotes
- Small RNA sequencing
- Single cell sequencing
- Whole genome resequencing
- 16S metagenome sequencing
- Meta-transcriptome sequencing after target depletion
- ChIP-sequencing
- Primary and secondary data analysis
- Bioinformatics services including data processing, data handling, and figure generation

PROTEOMICS/METABOLOMICS
The Proteomics/Metabolomics core facility at the Center for Biotechnology and Genomics provides services to the entire TTU community as well as outside customers.
We offer analysis of proteins and metabolites from cells, tissues, or other biological samples using cutting-edge equipment and methods.
Instrumentation:
- ABI/SCIEX 4800 MALDI-TOF/TOF mass spectrometer
- LTQ Velos Orbitrap nanospray mass spectrometer
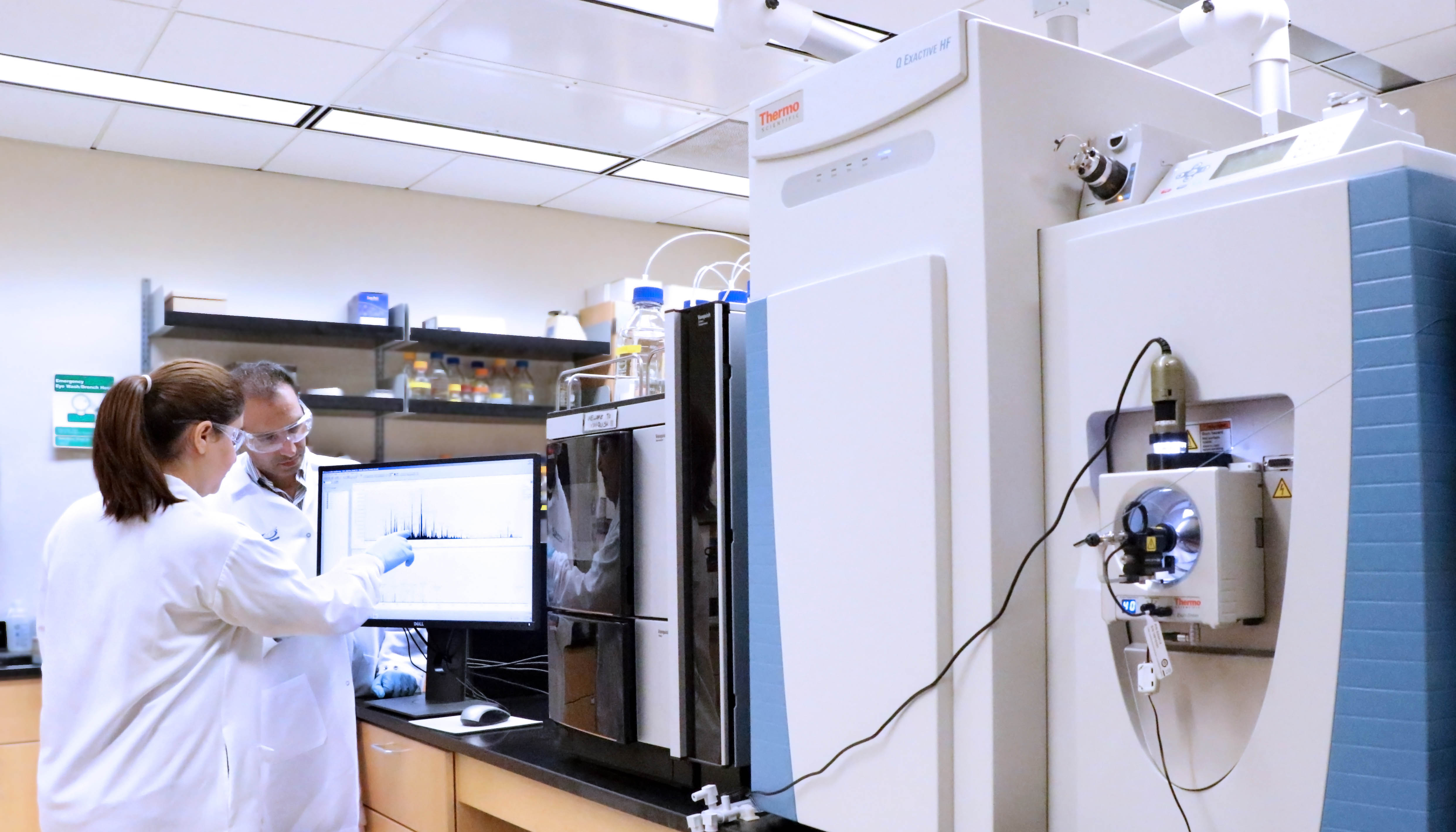
- Thermo Q-Exactive HF mass spectrometer
- Two Dionex-Ultimate 3000 Nano-LC systems
- Vanquish UHPLC
- Dionex-Ultimate 3000 with fraction collector for peptide fractionation and a UV/VIS detector
Services Offered:
- Protein IDs by LC-MS/MS and MALDI-TOF/TOF
- Whole proteome profiling from cells, tissue, etc.
- In-gel/in-solution digestion of proteins
- Data analysis using advanced software such as MASCOT, Proteome Discoverer, Compound Discoverer, LipidSearch, and Ingenuity Pathway Analysis (IPA)
- Protein quantification (label-free quantification, tagging)
- Analysis of post-translational modifications in proteins (e.g. phosphorylation)
- Global metabolite/lipid profiling
- Targeted metabolite analysis (identification and quantification)

BIOINFORMATICS
The Bioinformatics core facility at the Center for Biotechnology and Genomics provides services in the domains of genomics, proteomics, and general bioinformatics.
The CBG has the unique ability to generate, analyze, and integrate transcriptome and genome level data with proteomic and metabolomic data. The integration of these data with the Center's bioinformatic resources provides powerful discovery tools for understanding biochemical and signaling pathway responses to various challenges. We are excited to offer bioinformatics services at no additional charge to our customers, including data processing, data handling, figure generation, and any additional bioinformatics work that facilitates research publication and strengthen proposed projects.
Services Offered:
Nextgen Sequencing (NGS)
- RNA-Seq analysis
- Small RNA-Seq analysis
- de novo transcriptome assembly and annotation
- Whole genome assembly
- Metagenome and metatranscriptome analysis
- Pathway analysis
Proteomics
- Proteome analysis for identification of proteins
- Quantitative proteome analysis
- Pathway analysis
- Analysis of post-translational modifications
- Computational proteomics
- Protein modeling
- Protein-protein and protein-ligand docking
General Bioinformatics Software Development
Brochure for Genomics, Proteomics, and Bioinformatics Services Provided at the Center
Center for Biotechnology & Genomics
-
Address
Texas Tech University, Canton & Main Experimental Sciences Building, Room 101, Lubbock, TX 79409, Mail Stop 3132 -
Phone
806.742.6927 | Fax: 806.742.3788 -
Email
chiquito.crasto@ttu.edu
